7. Understanding Surfaces II
A quick note about the practicals...
Remember that coding should be done in Spyder using the understandinggis virtual environment. If you want to install the environment on your own computer, you can do so using the instructions here.
You do not have to run every bit of code in this document. Read through it, and have a go where you feel it would help your understanding. If I explicitly want you to do something, I will write an instruction that looks like this:
This is an instruction.
Remember that we are no longer using Git CMD. You should now use GitHub Desktop for version control.
Don't forget to set your Working Directory to the current week in Spyder!
Shortcuts: Part 1 Part 2
In which we learn to determine whether or not something is visible, as well as speed up our raster processing by replacing geometry with simple graphics algorithms.
Part 1
Last week we began our examination of raster data formats through the creation of a simple flood-fill algorithm. This week we will extend this and learn to calculate visibility - i.e., whether or not something is visible from a given location. Visibility analysis is extremely common in GIS, which makes it all the more surprising how little it is understood by GIS practitioners and decision makers!
One of the interesting things about visibility analysis in GIS is that there are a great deal of different approaches to it, with varying levels of accuracy and speed. A comparison of some common approaches is given by Kaučič and Zalik (2002). This week, we will be using a simplified version of my Viewshed algorithm, which is similar to the 8WS approach described in the paper.
One of the key features of this approach is that we will once again exploit working in image space in order to allow us to replace complex geometric calculations with simple graphics algorithms in order make our code more efficient (just as we did with the flood fill algorithm last time). Many of these algorithms were created in the early days of computer graphics, and so are designed to be extremely efficient (as computer power was so limited at the time).
Specifically, we will use two algorithms:
- Bresenham’s Line algorithm, which draws a line across a surface from one pixel to another.
- Midpoint Circle algorithm (a.k.a. Bresenham’s Circle Algorithm), which draws a circle of a given size (in pixels) around a given pixel.
These algorithms are extremely efficient because they allow us to draw lines and circles simply using the + and - operators, completely avoiding the use of trigonometry, which would be much more complex (see here to remind yourself of the equations used in trigonometry approaches).
We will not be implementing the Bresenham’s Line or Midpoint Circle algorithms directly - instead we will use the implementations that are provided by the draw module of the scikit-image library, which contains a range of functions for image processing in Python. This will allow us to focus on the details of the visibility calculation itself (which we cover in Part 2). However, if you want to know more about these algorithms, you can read about both of them in the original paper by Bresenham (1965).
Because we can apply algorithms like those provided in scikit-image, it is quite a good rule of thumb for raster-based algorithms that the more work you can do in Image Space, the faster your code could be. Also remember the golden rule to help avoid errors - try to work entirely in either image space or coordinate space - avoid mixing and matching if you can!
Take a look at the selection drawing algorithms available in
scikit-imagethey might come in handy in the future… (hint hint…)
CRS
As with last week, our data are already projected to the British National Grid (EPSG: 27700) - a local conformal projected CRS designed by Ordnance Survey for general use in Great Britain. For this reason, we do not need to select a CRS this week, as we know that this is an acceptable CRS for use in our area of interest (the Lake District).
Setting up a new Raster Layer
Last week, we learned how to open a raster layer and make a blank copy of it using the zeros() function from numpy. Let’s set one of those up now using the same dataset (Lake District, Cumbria, UK), just as we did last week:
from numpy import zeros
from rasterio import open as rio_open
from rasterio.plot import show as rio_show
from matplotlib.pyplot import subplots, savefig
from matplotlib.colors import LinearSegmentedColormap
# open the elevation data file
with rio_open("../data/helvellyn/Helvellyn-50.tif") as dem:
# read the data out of band 1 in the dataset
dem_data = # COMPLETE THIS LINE
# create a new 'band' of raster data the same size
output = # COMPLETE THIS LINE
# plot the dataset
fig, my_ax = subplots(1, 1, figsize=(16, 10))
# add the DEM
rio_show(
dem_data,
ax=my_ax,
transform = dem.transform,
)
# add the drawing layer
rio_show(
output,
ax=my_ax,
transform=dem.transform,
cmap = LinearSegmentedColormap.from_list('binary_viewshed', [(0, 0, 0, 0), (1, 0, 0, 0.5)], N=2)
)
savefig('./out/bresenham.png', bbox_inches='tight')
Create a new file called
bresenham.pyin yourweek8directory, add the above snippet and run it
Remember that we used LinearSegmentedColormap from the matplotlib.colors library to create our own colormap, in which values of 0 are set to transparent black (0, 0, 0, 0), and values of 1 are set to a different colour. Last week we chose a shade of blue (to represent flood water), this week we will use semi- red (1, 0, 0, 0.5)
Make sure that you understand the above (if not, refer back to the last practical!) and then run your code
If all goes to plan, you should have a map of the DEM (because the new image is all 0s, and so completely transparent):
You have added a new file to your repo, so you need to run git add week8/bresenham.py or git add . to make sure your new file is tracked by git. After that, it’s…

Drawing to your new Raster Layer
Drawing a Point
Now we have an transparent layer laying perfectly over our map, and we know that any pixels that we set to 1 will appear as red.
As with last week, let’s start by drawing a point. Remember that the steps for this are:
- Convert the coordinate pair from coordinate space (
x,y) to image space (row,col) (as with last week, you should create acoords_2_img()function). - Set the pixel at this location in your
outputsurface to1
Using the above and your knowledge from last week, see if you can draw a point at
334170, 515165. Though it is perfectly possible to do this in a single line of code, I want you to use two lines of code (one for each step above), storing the image space coordinates in variables calledrowandcol- this is because we are going to use these values again in a moment.
If all went to plan, you should see the summit of Helvellyn go red (as with before - it is one tiny pixel, so you need to look closely!):
Great! Now we have drawn a point, let’s try drawing a line…

Drawing a Line (using Bresenham’s Line Algorithm)
We could easily draw a line by working out which cells to colour using trigonometry, but we will be using the (much more efficient) Bresenham’s Line Algorithm. We will use it via the line() function from the draw module of scikit-image.
This is a simple function that gives us a list of pixel locations (given in image space) that describe a line between two locations (also given in image space). We will demonstrate this by drawing a 50 pixel line due east from our existing point location. line has 4 arguments: the first two are the coordinates of the cell in which the line should start, and the second two are the coordinates of the cell in which the line should finish (both coordinates are in image space, and so row, col).
Add the below snippets to your code and run it to see the results
from skimage.draw import line
print(line(row, col, row, col+50))
If all went to plan, you should see something like this (I have shortened this using ellipses ... - yours should have a tuple of two numpy arrays each containing 51 numbers):
(array([296, 296, 296, ..., 296, 296, 296]), array([283, 284, 285, ..., 331, 332, 333]))
numpy arrays (a type of Python collection provided by the numpy library) are similar to a list, but with more restrictions to facilitate vectorisation (e.g., all of the elements in the array must be of the same type, meaning that the size of the array in memory can be predicted, and so the elements accessed directly without a pointer).
Note the use of the column_stack() function from the numpy library, which simply ‘stacks’ the two one-dimensional arrays (one for row indices, and one for column indices) returned from line() into a two-dimensional array (with each element in the form (row, column)) vectorisation (e.g. they can only store data of a single type) and more member functions.
Here we have a tuple (another type of Python Collection, with which you are already familiar) of two arrays: one array contains the row coordinates, and the other contains the column coordinates. This is great, but would be more useful if the coordinates were paired up. Fortunately, numpy has us covered once again, with the column_stack() function, which simply ‘stacks’ the two one-dimensional arrays (one for row indices, and one for column indices) returned from line() into a two-dimensional array (with each element in the form (row, column)):
from numpy import column_stack
print(column_stack(line(row, col, row, col+50)))
Add the above
importstatement to your code (remembering that you can combine it with an existing one…), and replace your print statement with the above one.
You should now be looking at something like this (again I have shortened this using ellipses ... - yours should have a list of 50 lists each containing 2 numbers):
[[296 283]
[296 284]
[296 285]
...
[296 331]
[296 332]
[296 333]]
Much better! Now we have a single array in which each item is a pair of coordinates (in image space) describing a pixel that is part of the line.
Make sure that you understand what
column_stackdoes - if you are not sure, ask!
Now all that remains is to loop through each of these coordinate pairs and update the value at that location in the output surface to 1:
Loop through each coordinate pair (
r,c) returned by this statement and set the output to 1 at that location, run your code and see what the result is
If all has gone to plan, you should be looking at something like this:
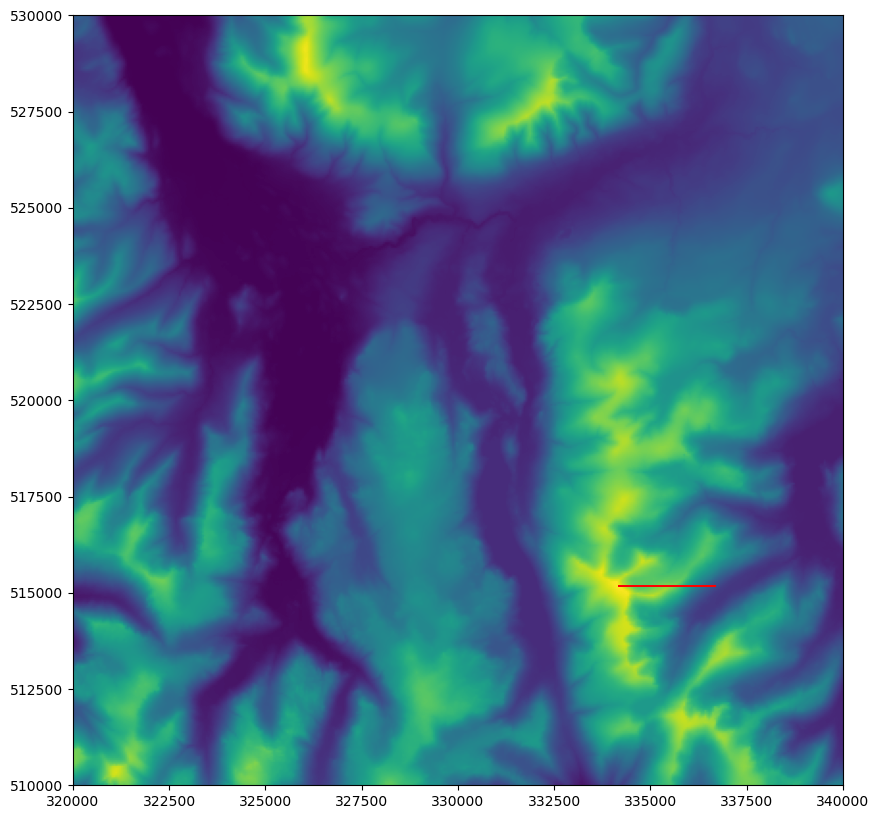
If so, then great!! On we go…

Drawing a Circle (using The Midpoint Circle Algorithm)
Finally, let’s add a circle in as well, we will draw one with a 50px radius around our point:
from skimage.draw import circle_perimeter
circle_perimeter(row, col, 50)
Implement the above snippets - you will need to pass the result to
column_stack(just as above) and set each cell in the resulting list to1Run your code
You should now be looking at something like this:
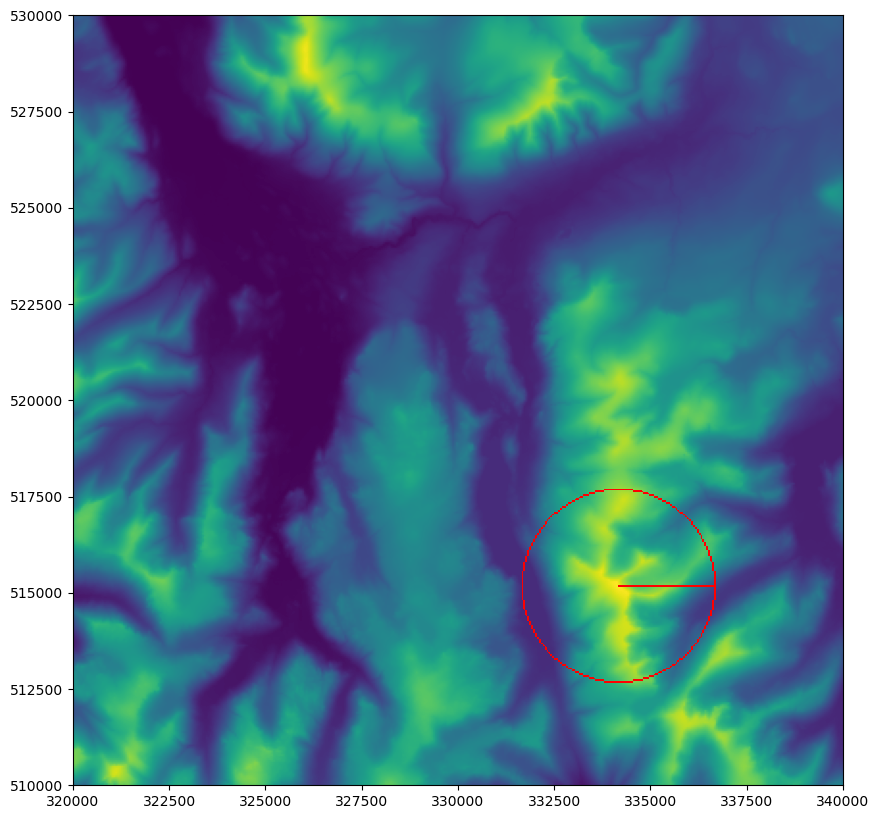
If so, well done! You can now successfully draw points, lines and circles to a raster - with no geometry in sight!

Part 2
Before we get stuck in with understanding viewsheds, let’s do a bit of setup:
Open
week8/week8.pyand add the necessary code to load"../../data/helvellyn/Helvellyn-50.tif"into a variable calleddemSet a location for the origin of the viewshed using the below snippet
# set origin for viewshed
x0, y0 = 330000, 512500
In a moment, we will create a viewshed() function. But first, we have a little more prep to do…
In order to allow us to test it, add the function call in the below snippet to your code:
# calculate the viewshed
output = viewshed(x0, y0, 20000, 1.8, 100, dem_data, dem.transform)
Note that, just as we did last week, we will need to pass both the data band (dem_data) and the transform object (dem.transform) to our viewshed() function. This is better than passing the DatasetReader object itself, as this is a large and complex object that can be rather unstable (because it points to a file, which could be closed or moved, for example).

Now we’re ready to start…
The viewshed() Function
As you saw in the lecture, our viewshed algorithm works by calculating a circle (using the Midpoint Circle Algorithm), and then calculating a line of sight to each location on the circle, updating each cell in the line as we go. Each of these two steps will be represented by a function - we will start by making the viewshed() function, and then move on to line_of_sight() later on.
Create a new function called
viewshed()with 7 arguments as listed below:
x0, the x coordinate (in coordinate space) of the point from which we will calculate the viewshedy0, the y coordinate (in coordinate space) of the point from which we will calculate the viewshedradius_m, the radius (in metres) of the viewshedobserver_height, the height (in metres) of the observer for whom the viewshed will be calculatedtarget_height, the height (in metres) of the target (e.g. a wind turbine) to which the viewshed will be calculateddem_data, the DEM data (the band from therasteriodataset)transform, an instance of theAffineclass that can be used to convert between coordinate space and image space
The viewshed() function will work out the endpoint for each line of sight (each cell in the circle) and then call the line_of_sight() function to work out the visibility of each point on a line from the origin to that location.
The first thing that it will do, however, is to convert everything from coordinate space to image space - it is important to do this straight away, as otherwise you can get mixed up, which is a common way to introduce errors into your code when working with rasters!
Based on your knowledge from last week, convert
x0andy0to image space, storing the results asr0andc0for the row and column value respectively.
Now that we have this - we have the opportunity to make our code more robust, by checking that the specified location is actually within the area covered by the dataset. We could have done this using the bounding box of the raster dataset and a within query using the coordinate space coordinates, but as above it is much simpler to do this in image space, and simply use the between operator to see if the coordinates are larger than 0 and smaller than the width/height of the dataset. If this is not the case (note how we are using the not operator here to invert the results of the conditional statement) we simply print a message to the user and gracefully exit the program.
from sys import exit
# make sure that we are within the dataset
if not 0 <= r0 < dem_data.shape[0] or not 0 <= c0 < dem_data.shape[1]:
print(f"Sorry, {x0, y0} is not within the elevation dataset.")
exit()
Make sure that you understand how the above statement works - particularly how the
.shapeproperty returns the height and width of the dataset, and then add the above snippets to your code at the appropriate locations
One snag when working in image space is that we often need to know the size of each cell, in order to be able to convert it to a distance. For example, the 50 pixel line that we drew above is rather meaningless, unless we know that each cell represents 50m on the ground, meaning that the line is
\[50_{cells} \times 50 m = 2500 m\]Fortunately, we can access the resolution of the dataset from the Affine Transformation Matrix stored in the transform variable. The matrix contains a separate value for the x and y resolution, but GIS datasets generally have the same resolution for both directions (i.e. the cells are square), so we can simply take one value for both, which is stored in transform[0].
Add the below snippet to convert the
radius_mto image space (from metres to pixels) by dividing it by the resolution of the dataset
# convert the radius (m) to pixels
radius_px = int(radius_m / transform[0])
Now we know the starting location and radius of the viewshed, we are just missing one key bit of information: the starting height of the viewshed - by which I mean the elevation of the origin location plus the observer height. To get this, we simply need to read the elevation from the DEM and add it to the observer height.
Complete the below snippet to calculate the sum of the elevation at
(r0, c0)and theobserver_height
# get the observer height (above sea level)
height0 =
Add a
print()statement to see what the elevation is at our chosen start point, run the code to get the result
(it should be about 608.1m).

Now we are nearly ready to start our viewshed:
Based on your knowledge from last week, use
zeros()to create an output raster surface at the same dimensions as thedemdataset, save it in a variable calledoutputSet the value of the viewshed origin (
(r0, c0)) to1in theoutputsurface (as the observer can definitely see the cell in which they are stood)
All we need to do now is calculate our circle, and pass each cell to the line_of_sight()function, before returning our output dataset to be added to the map:
# get pixels in the perimeter of the viewshed
for r, c in column_stack(circle_perimeter(r0, c0, radius_px)):
# calculate line of sight to each pixel, pass output and get a new one back each time
#output = line_of_sight(r0, c0, height0, r, c, target_height, radius_px, dem_data, transform, output)
pass
# return the resulting viewshed
return output
The above snippet should be relatively self explanatory - note (as above) the use of the column_stack() function to convert the index values into a single array of coordinate pairs.
As you can see, each call provides all of the necessary arguments to line_of_sight(), including the current version of output , which is then replaced by an updated version after the line_of_sight() function has set all of the visible cells on that line to 1. The effect is that output is overwritten with a more complete version every iteration of the loop, until every line has been assessed and the viewshed is complete.
Also note the use of pass here - this is a temporary placeholder Python statement that does nothing, but avoids the error that would arise from having an empty block in the for loop while the call to the line_of_sight function is commented out.
Make sure that you understand the above snippet, and add it to your code
Check that your code runs without error. If so, un-comment the line that begins
output = ...from the above snippet and delete thepassstatement.

Now we can get down to business with our calculation…
The line_of_sight() Function
As you will remember from the lecture, as we move along a line of sight, we check each cell in turn, and work out the \(\Delta y /\Delta x\). If it is bigger than any we have seen previously, then the cell is visible. If it is not bigger, then the cell is invisible - simple!
Create a new function called
line_of_sight()with 10 arguments as listed below:
r0, therowcoordinate of the pixel at the start of the line (in image space)c0, thecolumncoordinate of the pixel at the start of the line (in image space)height0, elevation+height of the pixel at the start of the line (in metres)r1, therowcoordinate of the pixel at the end of the line (in image space)c1, thecolumncoordinate of the pixel at the end of the line (in image space)height1, the height of the pixel at the end of the lineradius, the radius of the viewshed (in pixels - image space)dem_data, the DEM data (the band from therasteriodataset)transform, an instance of theAffineclass that can be used to convert between coordinate space and image spaceoutput, the output dataset to which the new data will be added
Our approach will be to loop along the line from (r0,c0) to (r1,c1), and calculate \(\Delta y /\Delta x\) at each cell. If it is greater than any previous \(\Delta y /\Delta x\) value, then the cell is visible, and so we set the value to 1 in the output dataset. If not, then the cell will remain at 0.
To get started, we will initialise a variable to hold the largest \(\Delta y /\Delta x\) so far, and then we will loop through each cell in our line (ignoring the first one, which we know is visible because our observer is stood in it…). Because the value could be positive or negative (depending on whether we are looking uphill or downhill), we will initialise our tracking variable to negative infinity (a special value that is guaranteed to be smaller than all of the other numbers Python can hold):
# init variable for biggest dydx so far (starts at -infinity)
max_dydx = -float('inf')
# loop along the pixels in the line (excluding the first one)
for r, c in column_stack(line(r0, c0, r1, c1))[1:]:
Note the use of the column_stack() function to convert the index values into a single array of coordinate pairs and the use of list slicing to excluse the first element from the loop (see the Hints Page if you can’t remember how this works).
Add the above snippet to your code
Inside the loop, we will firstly calculate \(\Delta x\):
Use Pythagoras’ Theorem (either implement yourself or use the
math.hypotfunction) to work out the distance (in pixels) between(r0,c0)(the origin point) and(r,c)(the current point) - store the result in a variable calleddxAdd the below snippet to stop the loop when it gets to the end of the desired radius (this is necessary because we used a larger radius to calculate the endpoint in order to increase the line density) OR when we hit the end of the dataset (as going past this would give us an error)
# if we go too far, or go off the edge of the data, stop looping
if dx > radius or not 0 <= r < dem_data.shape[0] or not 0 <= c < dem_data.shape[1]:
break
Temporarily add a
print()statement so that you can see the distances - remember to remove this once you have verified that it works!
Next, we just need to work out the \(\Delta y /\Delta x\) value - which we do twice (once for the base of the object that we are looking for, and once for the tip). This is because, when assessing the visibility of the object for this cell, we need to use the tip (as the object will actually be located there); but when we are testing the visibility of the object further down the line, then we want to account for the ground level at that location (as the object will not be there to block visibility).
# calculate the current value for dy / dx
base_dydx = (dem_data[(r, c)] - height0) / dx
tip_dydx = (dem_data[(r, c)] + height1 - height0) / dx
Make sure that you understand that (if not, ask!) and add the above snippet to your code
Make sure that it runs without error
Finally, we just need to test if this \(\Delta y /\Delta x\) was the biggest so far, and if so update the cell in the output surface accordingly. We can then return the freshly edited output to the viewshed() function, which will add the rest of the lines of sight to it, before returning it to our main code block where we can draw the result!
Add the below snippet to your code to complete the
line_of_sight()function:
# if the tip dydx is bigger than the previous max, it is visible
if (tip_dydx > max_dydx):
output[(r, c)] = 1
# if the base dydx is bigger than the previous max, update
max_dydx = max(max_dydx, base_dydx)
# return updated output surface
return output
Make sure that it runs without error

Drawing your map
If that all runs without error - then there is a very good chance that you are successfully calculating a viewshed! To check - we can draw it to a map - there is nothing new here (we did it all last week or in Part 1), so I will give this in a single block (I will stick with the default 'viridis' colour ramp, but feel free to change it to another one!):
from math import floor, ceil
from geopandas import GeoSeries
from shapely.geometry import Point
from matplotlib.lines import Line2D
from matplotlib.patches import Patch
from matplotlib.colors import Normalize
from matplotlib.cm import ScalarMappable
from rasterio.plot import show as rio_show
from matplotlib.pyplot import subplots, savefig
from matplotlib_scalebar.scalebar import ScaleBar
from matplotlib.colors import LinearSegmentedColormap
# output image
fig, my_ax = subplots(1, 1, figsize=(16, 10))
my_ax.set_title("Viewshed Analysis")
# draw dem
rio_show(
dem_data,
ax=my_ax,
transform = dem.transform,
cmap = 'viridis',
)
# draw dem as contours
rio_show(
dem_data,
ax=my_ax,
contour=True,
transform = dem.transform,
colors = ['white'],
linewidths = [0.5],
)
# add viewshed
rio_show(
output,
ax=my_ax,
transform=dem.transform,
cmap = LinearSegmentedColormap.from_list('binary_viewshed', [(0, 0, 0, 0), (1, 0, 0, 0.5)], N=2)
)
# add origin point
GeoSeries(Point(x0, y0)).plot(
ax = my_ax,
markersize = 60,
color = 'black',
edgecolor = 'white'
)
# add a colour bar
fig.colorbar(ScalarMappable(norm=Normalize(vmin=floor(dem_data.min()), vmax=ceil(dem_data.max())), cmap='viridis'), ax=my_ax, pad=0.01)
# add north arrow
x, y, arrow_length = 0.97, 0.99, 0.1
my_ax.annotate('N', xy=(x, y), xytext=(x, y-arrow_length),
arrowprops=dict(facecolor='black', width=5, headwidth=15),
ha='center', va='center', fontsize=20, xycoords=my_ax.transAxes)
# add scalebar
my_ax.add_artist(ScaleBar(dx=1, units="m", location="lower right"))
# add legend for point
my_ax.legend(
handles=[
Patch(facecolor=(1, 0, 0, 0.5), edgecolor=None, label=f'Visible Area'),
Line2D([0], [0], marker='o', color=(1,1,1,0), label='Viewshed Origin', markerfacecolor='black', markersize=8)
], loc='lower left')
# save the result
savefig('./out/7.png', bbox_inches='tight')
print("done!")
Add the above snippet to your code at the appropriate location and run it
If all has gone to plan, you should have something like this:
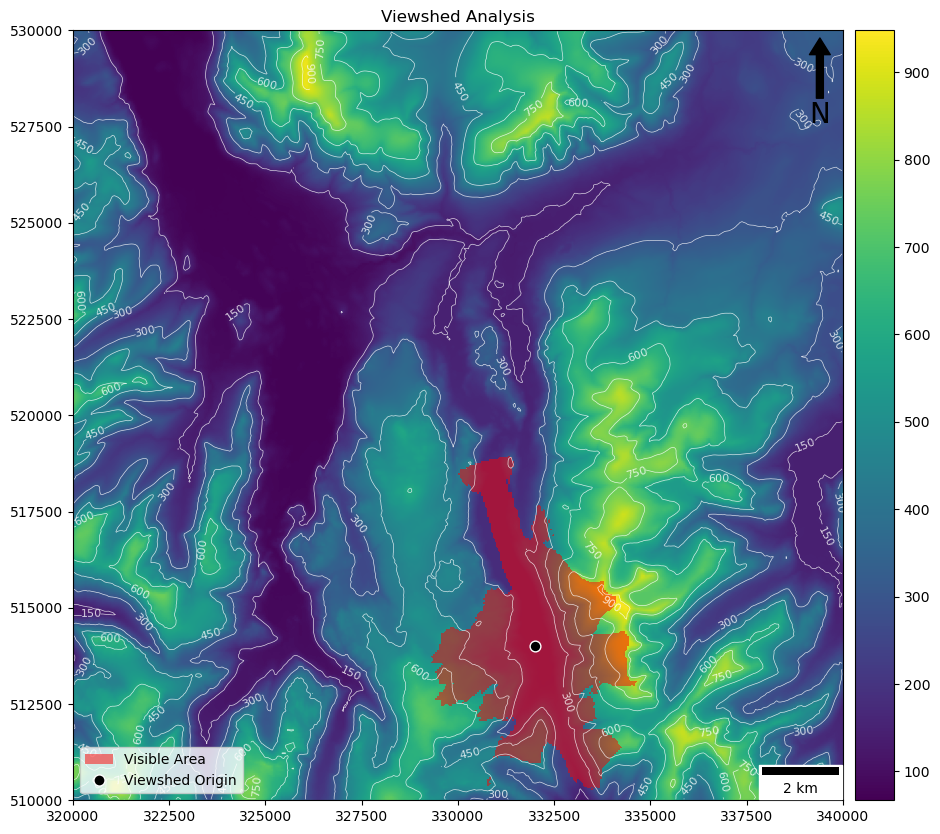
(where the red represents the area visible from the black dot). If you think about it (using the colour ramp and contours to help), it should be clear to you why you can’t see the areas that are not included (for example, why you can’t see into the valley to the west)
If so then congratulations - you have calculated your own viewshed!!

Try adjusting your origin point and radius (coordinates are shown on the axes) to see if the pattern of visibility matches with your expectations. You will notice how the chosen radius is extremely important, and that increasing the radius affects the speed of execution.
A little more…
If you want to go a little further, we can add one more function to account for atmospheric refraction and the curvature of the Earth. This can be achieved with a relatively simple equation:
\[height_{adjusted} = height - \frac{\Delta x^{2}}{D} \times (1 - R)\]where:
\(height_{adjusted}\) is the resulting (adjusted) height
\(height\) is the original (un-adjusted) height
\(\Delta x\) is the distance in metres
\(D\) is the diameter of the Earth in metres
\(R\) is the atmospheric refraction coefficient
Add a new function using the snippet below, and complete it to reflect the above equation
Edit your
base_dydxandtip_dydxlines inline_of_sight()to make use of this function
def adjust_height(height, distance, earth_diameter=12740000, refraction_coefficient=0.13):
"""
* Adjust the apparant height of an object at a certain distance, accounting for the
* curvature of the earth and atmospheric refraction
"""
return
You will note that we have used some optional arguments in this function (earth_diameter=12740000, refraction_coefficient=0.13). When I do this, I can either choose to define these values or just ignore them and accept the default values. For example, I could set all of the values as normal:
adjust_height(height, distance, 12800000, 0.15)
…or I can just ignore them and accept the defaults:
adjust_height(height, distance)
…or I can use a named argument and just set one of them, and accept the default for the other (which allows for them not to be in the right order because I have missed one):
adjust_height(height, distance, refraction_coefficient=0.15)
You will need to have a think about how to calculate the distance value (in metres) too - remember you already have a value for dx (the distance in pixels) and you know the resolution (size of a pixel), so it should be easy enough to work out…